SomaScan®
Proteomics Consortium
Advancing precision medicine research through worldwide sharing of human proteomic data
Advancing the power of precision medicine and population proteomics research through global collaboration and data sharing
The SomaScan Proteomics Consortium was founded by Russell Bowler, MD, PhD, to enable researchers to increase the power of their studies by leveraging high-plex proteomic profiling with the SomaScan® Assay across diverse populations worldwide, for more powerful biomarker discovery and deeper insights into human biology. Members are committed to sharing data for the purpose of increasing efficiency and efficacy, as well as for fostering discussions about the development and pursuit of new studies and research directions.
SomaLogic is honored to support the consortium as an independent and open exchange of ideas, knowledge, and results, in order to accelerate the study of disease and human biology.
For general inquiries or to join the consortium, please contact Russell Bowler, MD, PhD.
Consortium Principles
- Advance knowledge of the proteome to improve understanding of disease etiology, diagnosis and prognosis
- Promote opportunities to collaborate and share results
- Adopt a distributive analysis model
- Preserve governance of cohort data with cohort Principal Investigators
- Support the development of early-career researchers
- Raise common funds through grants and donations by sharing analyses and preliminary data
Current cohorts used by consortium members in proteomics research using the SomaScan Assay
To learn more about the biobanks and prospective cohort studies listed, please click “View now” next to each.
Data tips and resources
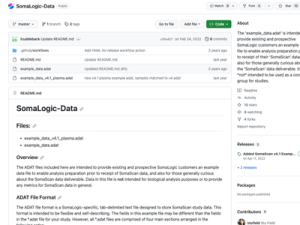
The ADAT file format is a Soma Logic-specific, tab-delimited text file designed to store Somascan study data. This format is intended to be flexible and self-explanatory. When sample analysis is completed, SomaLogic normalizes the data to ensure data quality and then creates an ADAT file for delivery of results to the researcher.
If you have questions or need assistance with ADAT files, please email SomaLogic Technical Support here.

A data repository serves as a structured digital storage facility, often resembling databases or data lakes, where research data and analytical resources are systematically organized and made accessible in accordance with FAIR (Findable, Accessible, Interoperable, and Reusable) standards. Data repositories exist in two primary forms: open access and closed access.
Learn why researchers are increasingly encouraged – or required – to upload data to a repository before publishing their findings.
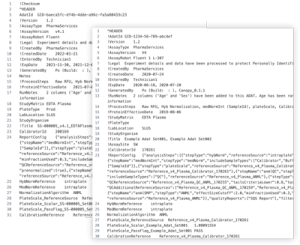
Data repositories can exist in two primary forms: open access, which are open to the public, or closed access, restricted to specific users or groups, and they may cater to discipline-specific or broader research datasets. Choosing which type of data repository to upload to depends on several factors. See the table below for details:
Researchers can analyze and visualize all their studies in one convenient view on the SomaLogic Life Sciences Portal. Digital resources, such as DataDelve™ Statistics and the SomaScan Menu, are easily accessed from the portal, too.
The SomaLogic Technical Support team is always available to support researchers and repository owners. Email us with questions.

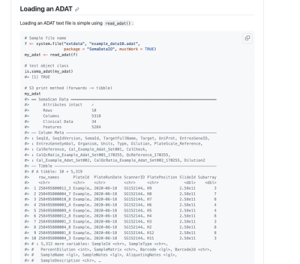
SomaScan data are presented in an ADAT format, a SomaLogic-specific, tab-delimited text file that’s small enough to fit on a standard desktop computer. ADAT files can be read easily into any R/Python scripting languages using our ADAT reader function in SomaDataIO (R package) or Canopy (Python package). For uploading to public data repositories, we recommend that SomaScan data owners provide ADAT files formatted as they received them from SomaLogic.
For more information on ADAT format or handling SomaScan data, please email the SomaLogic Technical Support team.
![]()
The SomaLogic Technical Support team is available throughout the analysis process – from initial questions and sample prep, to bioinformatics analysis and questions and final delivery of data.
Please email our team here.
Deepen your molecular insights with the SomaScan Platform
The SomaScan Platform is a single platform from discovery to development that gives researchers access to half the genetically encoded human proteome (11,000 protein measurements). The technology is powered by SOMAmer® (Slow Off-rate Modified Aptamer) Reagents, which provide researchers with reliability and reproducibility unparalleled in the proteomics industry.
Read more at the links below:




