Plasma Proteomics 101: Why It’s Central to Biomarker Discovery
When it comes to studying health and disease, plasma is the gold standard. It’s the most commonly collected biological sample in medicine, with hundreds of millions of tubes drawn every year for diagnosis and monitoring1. But why is plasma the go-to choice for biomarker discovery researchers?
Easy to access, rich in biological insight
One reason is simple practicality: plasma is easy, safe and minimally invasive to collect through a routine blood draw. This accessibility supports repeated sampling over time, making plasma well suited for longitudinal studies that track disease progression and treatment response.
By comparison, more invasive sampling methods – such as extracting cerebrospinal fluid (CSF) or synovial fluid – can be painful and carry greater risks of complications. Other sample types like saliva and skin can offer useful clues into health and disease, but typically reflect localized biology rather than system-wide processes1,2.
What is plasma proteomics?
Plasma proteomics is the systematic measurement and analysis of proteins circulating in plasma or serum, the liquid portion of blood after cells are removed3. The distinction between plasma and serum is technical: serum is collected after blood clots, while plasma is collected with anticoagulants and retains clotting proteins (Figure 1).
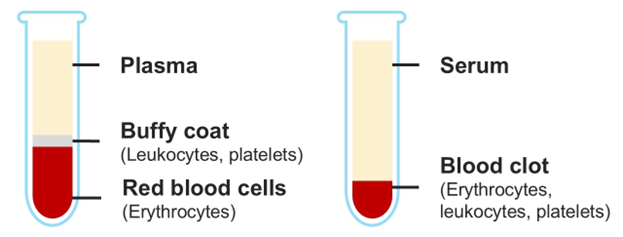
Figure 1. Comparison of plasma and serum. To prepare plasma (left tube), blood is collected in tubes containing anticoagulants and then centrifuged. This separates the blood into three components: plasma (top layer), the buffy coat (leukocytes and platelets), and red blood cells (bottom layer). To prepare serum (right tube), blood is collected without anticoagulants and allowed to clot before centrifugation. The clot (containing red and white blood cells and platelets) is separated from the serum (top layer). In both cases, the plasma or serum is carefully transferred to a new tube for analysis.
In practice, both plasma and serum are used in biomarker discovery. For the purposes of this blog, “plasma” is used as an umbrella term to represent plasma- or serum-based proteomic samples.
What makes up the plasma proteome?
Plasma contains thousands of proteins that together offer a real-time snapshot of what’s happening inside the body. This is supported by approximately 4 liters (1 gallon) of lymph entering the bloodstream each day, delivering proteins from a broad spectrum of organs and tissues4. These include:
- Proteins secreted by healthy or diseased tissues
- Hormones and signaling molecules like cytokines
- Immunoglobulins (antibodies)
- Proteins leaked from damaged or dying cells
- Foreign proteins from pathogens
A window into health and disease
Changes in plasma protein levels have been linked to conditions ranging from head injuries and meningitis to sepsis and normal developmental changes1,5,6. These signals, found in just a few drops of blood, can open the door to earlier diagnosis, better treatments and a deeper understanding of human health.
During disease, certain proteins may enter the bloodstream, sometimes long before symptoms appear. Many of these proteins are present at very low levels or undetectable in healthy individuals, but become elevated during pathological processes, such as acute injury, cancer progression or neurodegeneration1,7. Here are some examples:
- Cardiovascular Disease: Cardiac troponins (cTnI, cTnT), CK-MB, Cardiac myoglobin
- Neurological Disorders: GFAP, Neurofilament light chain (NfL), Tau protein
- Cancer: Tumor-associated antigens (e.g., CA-125, PSA, CEA), Aberrantly glycosylated proteins and oncoproteins
- Infection & Sepsis: Procalcitonin (PCT), Pathogen-derived proteins
- Liver Disease: ALT, AST, alpha-fetoprotein (AFP)
Despite its value, plasma is also one of the most analytically difficult biological samples to study in proteomics.
References
- Anderson, N Leigh, and Norman G Anderson. “The human plasma proteome: history, character, and diagnostic prospects.” Molecular & cellular proteomics : MCP 1,11 (2002): 845-67.
- Malmström, Erik et al. “Human proteome distribution atlas for tissue-specific plasma proteome dynamics.” Cell 188,10 (2025): 2810-2822.e16.
- Sapan, C. et al. “Considerations Regarding the Use of Blood Samples in the Proteomic Identification of Biomarkers for Cancer Diagnosis.” Cancer genomics & proteomics 3,3-4 (2006): 227-230.
- Moore, J.E. et al. “Lymphatic System Flows.” Annual review of fluid mechanics 50 (2018): 459-482.
- Deutsch, E.W et al. “Advances and Utility of the Human Plasma Proteome.” Journal of proteome research 20,12 (2021): 5241-5263.
- Bhowmick, P. et al. “An Update on MRMAssayDB: A Comprehensive Resource for Targeted Proteomics Assays in the Community.” Journal of proteome research 20,4 (2021): 2105-2115.
- Rice, S.J. et al. “Characterization of effective, simple, and low-cost precipitation methods for depleting abundant plasma proteins to enhance the depth and breadth of plasma proteomics.” Proteomics 24,15 (2024): e2400071.
More blogs
BlogAntibodies: The Hidden 20% of the Plasma Proteome
Antibodies make up 20% of plasma yet most proteomics studies overlook their specificity and early disease signals.
BlogHigh-Plex Affinity Proteomics for Plasma
High-plex affinity proteomics enables deep, reproducible plasma biomarker discovery. See how the SomaScan 11K Assay delivers scale, sensitivity, and coverage.
BlogThe Challenges of Plasma Proteomics for Biomarker Discovery
Why is protein biomarker discovery in plasma so difficult? Explore how protein complexity and dynamic range limit detection of low-abundance signals.





