High-Plex Affinity Proteomics for Plasma
High-plex affinity-based proteomics facilitates the detection of proteins across a broad concentration range. Among these approaches, the SomaScan™ 11K Assay is one of the most extensively validated platforms for plasma proteomics, with demonstrated performance across discovery, translational research and population-scale studies.
Designed for depth, scale and reproducibility
The SomaScan 11K Assay enables broad plasma proteome coverage without altering standard workflows or throughput, supporting large-scale and longitudinal study designs (Figure 1)1-3.
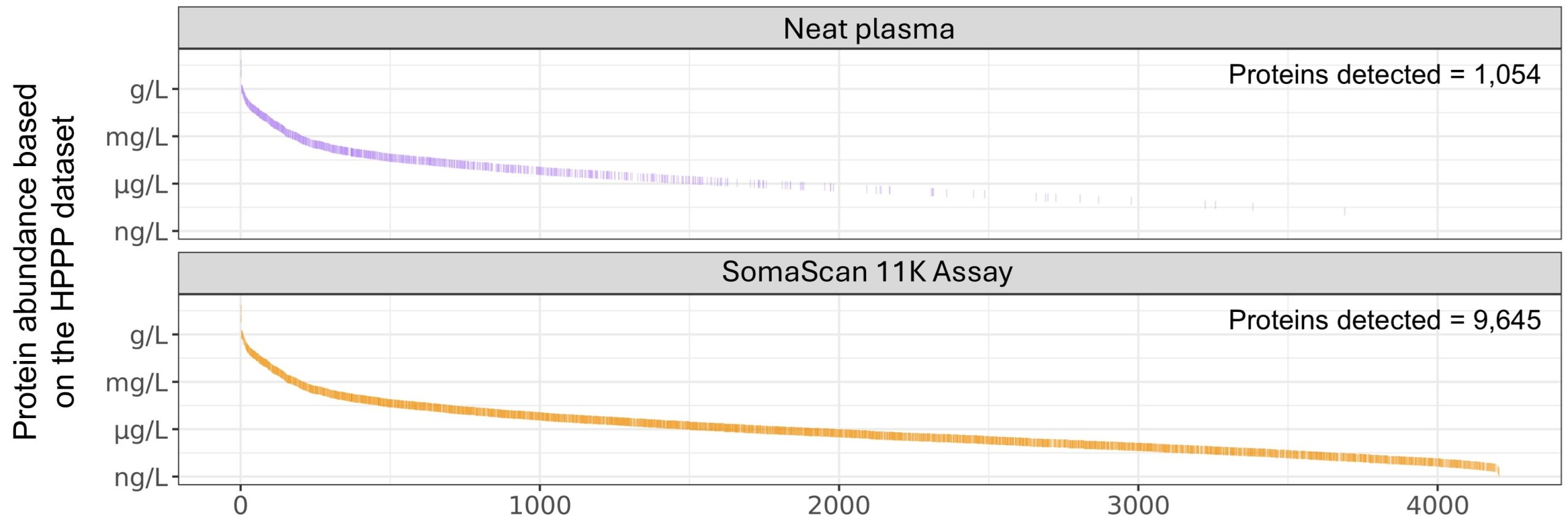
Figure 1. The SomaScan 11K Assay measures proteins across the full dynamic range of the plasma proteome. Comparison of proteins detected in neat plasma by mass spectrometry (purple) or the SomaScan 11K Assay (orange). Only proteins that overlap with the Human Plasma Proteome Project (HPPP) dataset are shown3. Data adapted from Beimers et al.1 and Kirsher et al.2.
Key features include:
- Broad coverage: Profile 11,000 protein measurements with high specificity and affinity, without depleting, enriching or fractionating plasma
- High sensitivity: Detect proteins at femtomolar (fM) levels directly in plasma
- Unmatched reproducibility: Median coefficient of variation (CV) of ~5%
- Wide dynamic range: Detect proteins across 10 logs of concentration
- Low sample volume: Requires only 55 µL of plasma per run
- No changes to existing workflows or throughput: Enables easy adoption in high-throughput settings
These features enable robust measurement of low-abundance signaling and immune-associated proteins, such as cytokines, chemokines and growth factors, that are central to disease mechanisms and therapeutic response.
Highest coverage of FDA-approved biomarkers
In an independent comparison of mass spectrometry workflows and affinity-based proteomics platforms, coverage of U.S. Food and Drug Administration (FDA)-approved protein biomarkers in plasma ranged from 43% to 89% (Figure 2)2,4. The SomaScan 11K Assay demonstrated the highest coverage across the eight platforms tested, measuring 89% of the biomarkers (193 of 217)2.
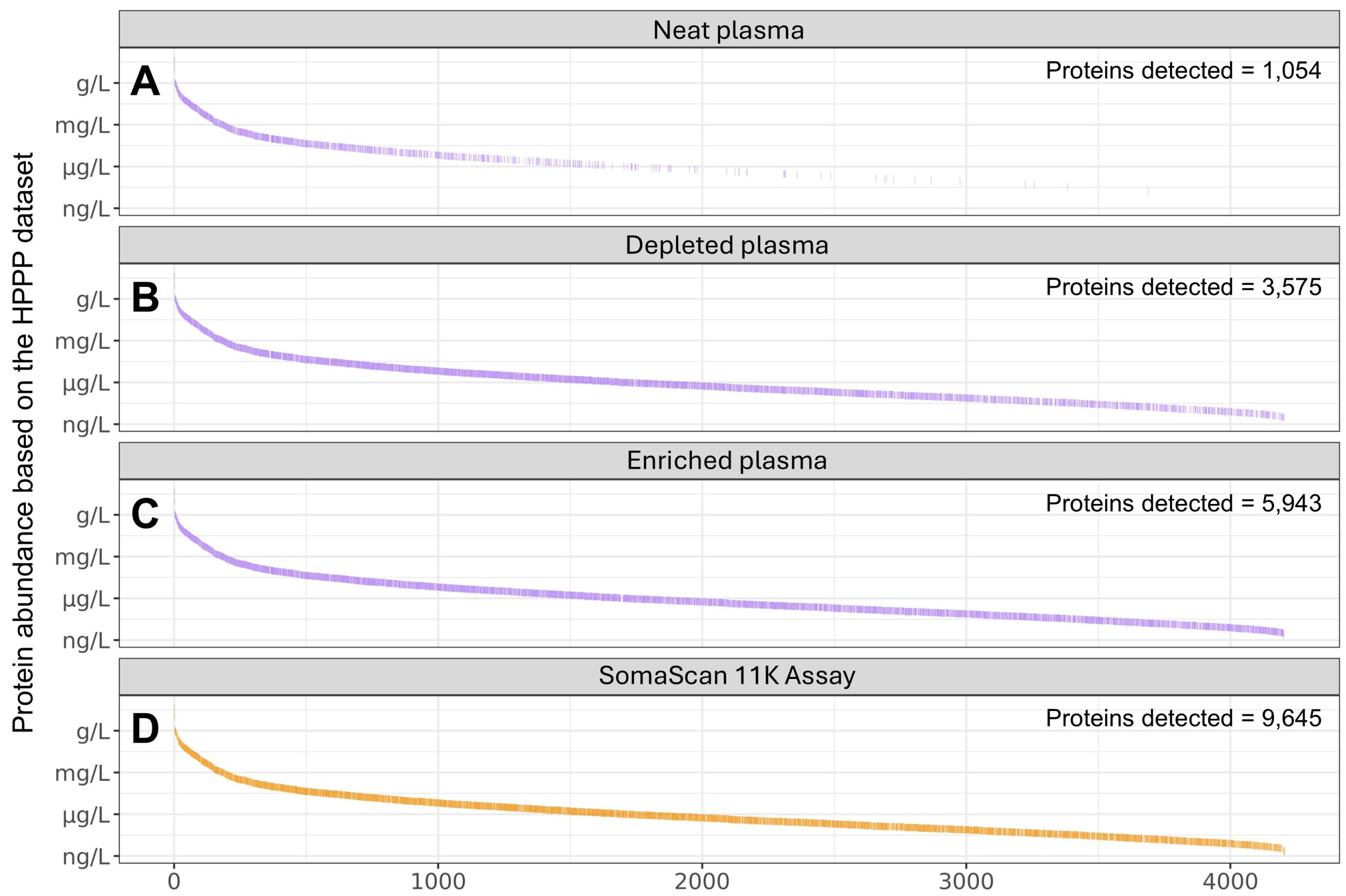
Figure 2. FDA-approved protein biomarkers detected in plasma by mass spectrometry and affinity-based proteomics. Proteins detected using mass spectrometry (purple) include those measured in: A) neat plasma, B) plasma with the top 14 most abundant proteins depleted, and C) enriched plasma. D) Proteins in neat plasma measured using an affinity-based proteomics platform, the SomaScan 11K Assay (orange). Detected proteins were compared against the list of FDA-approved plasma protein biomarkers reported by Bhowmick et al. Data for panel A were adapted from Beimers et al.1, and data for panels B–D were adapted from Kirsher et al.2.
Proven in population-scale studies
High-plex plasma proteomics must perform reliably not only in small discovery cohorts, but also across thousands of samples. The SomaScan Assay has been selected for use in approximately 270,000 plasma samples across diverse population-scale studies, including:
- Population health biobanks
- Cardiovascular and metabolic disease cohorts
- Neurodegeneration and aging studies
- Immunology and inflammation research
- Precision medicine and multi-omics initiatives
These efforts demonstrate that deep plasma proteomics can be performed reproducibly at scale, enabling analyses that move beyond single biomarkers toward integrated, systems-level insight across populations.
From plasma complexity to actionable insight
By combining high sensitivity, broad coverage and specificity, the SomaScan 11K Assay makes it possible to translate plasma complexity into biologically meaningful and clinically relevant insight across discovery, translational and population-scale research.
References
- Kirsher, D.Y. et al. “Current landscape of plasma proteomics from technical innovations to biological insights and biomarker discovery.” Communications chemistry 8,1 279. 25 Sep. 2025.
- Beimers, W.F. et al. “Technical Evaluation of Plasma Proteomics Technologies.” Journal of proteome research 24,6 (2025): 3074-3087.
- Deutsch, E.W. et al. “Human Proteome Project Mass Spectrometry Data Interpretation Guidelines 3.0.” Journal of proteome research 18,12 (2019): 4108-4116.
- Bhowmick, P. et al. “An Update on MRMAssayDB: A Comprehensive Resource for Targeted Proteomics Assays in the Community.” Journal of proteome research 20,4 (2021): 2105-2115.
More blogs
BlogAntibodies: The Hidden 20% of the Plasma Proteome
Antibodies make up 20% of plasma yet most proteomics studies overlook their specificity and early disease signals.
BlogThe Challenges of Plasma Proteomics for Biomarker Discovery
Why is protein biomarker discovery in plasma so difficult? Explore how protein complexity and dynamic range limit detection of low-abundance signals.
BlogPlasma Proteomics 101: Why It’s Central to Biomarker Discovery
Plasma proteomics is central to biomarker discovery. Understand the plasma proteome, its complexity, and how blood proteins reveal health and disease.





